Monster TV
Polecamy w Monster TV
Artykuły tylko dla bojowników JM (+18)
Zobacz galerię
Kto jest online?
Ale zawsze możesz się zarejestrować.
Gościmy obecnie 7302 osób, w tym 1097 rejestrowych bojowników walki z powagą.
Ostatnio dołączył do nas bojownik nagdennn
Faktopedia – Niewidzialny atrament brytyjskiej Secret Service
Dziś m.in. niewidzialny atrament brytyjskiej Secret Service, ptak odesłany do USA pierwszą klasą oraz podróżnik-twardziel ze Szwecji.


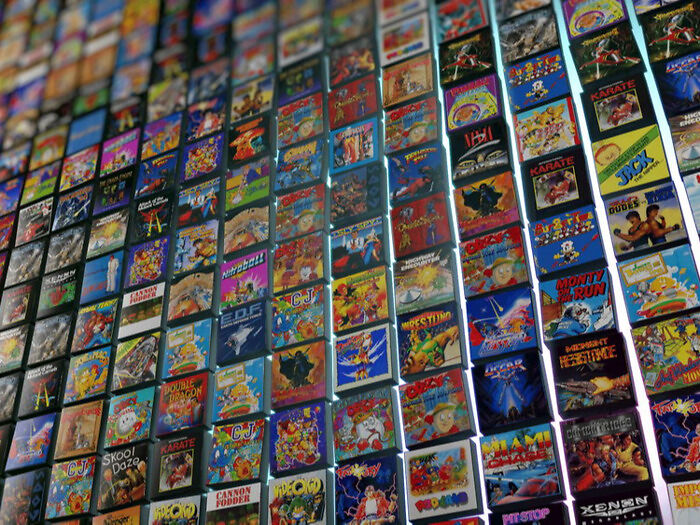
Będę grał w grę, ale jaką? Rozpoznasz grę po obrazku? Kultowe gry 8-bitowe. Boomerzy na start!

13 075
138
Dla fanów naprawdę klasycznych gier i grania. Poniżej screeny z różnych bardzo znanych czy wręcz kultowych gier. Ile z nich rozpoznasz? I zgadujemy PRAWIDŁOWE tytuły gier i na to pragnę zwrócić uwagę, gdyż w paru przypadkach te gry były piracone pod innymi tytułami.

Największe obciachy – Tego na komuniach jeszcze nie było
Dzisiaj:

- Tak się zdenerwował na krany, że je wyciął
- Młotek i lampa, czyli jak nie wymieniać żarówki
- Trzy miesiące więzienia dla byłego polityka PiS za zabicie psa
- Poszukiwacze taksówek nad Morskim Okiem

Dzięki temu gadżetowi idiotów będzie można poznać z daleka – Demotywatory

- Ruszyła pierwsza edycja konkursu Miss AI
- Ile samochodów policja skonfiskowała pijanym kierowcom do tej pory?
- Odbył się pierwszy Kongres Mężczyzn – podpiszecie się pod jego postulatami?
- Sekret kart telefonicznych TP SA

Ten film bije rekordy popularności na TikToku i nikt nie wie, dlaczego tak jest – Co nowego w technologii?
Dzisiaj:

- Brazylia pozywa Elona Muska
- Zakaz TikToka w USA już praktycznie sfinalizowany. Amerykanie żegnają się z platformą
- Microsoft zaprezentował nowy model AI i wygląda to dość niepokojąco
- W sieci pojawił się film pokazujący, gdzie i kiedy latała Taylor Swift w 2023 roku

Krótki dżołk
Nie udał się strajk listonoszy Poczty Polskiej. Okazuje się, że zamiast postulatów zostawili awizo.
W pracy znalazłem jakąś limitowaną holograficzną monetę – Bardzo nietypowe znaleziska internautów
Czasem w życiu natrafiamy na coś, co potrafi wzbudzić u nas na tyle duże zainteresowanie, że aż musimy się tym podzielić ze światem.


Jak rozpoznać 15 popularnych formacji chmur?
Wszyscy znamy cumulusa. Cumulus to archetypowa chmura: puszysta i biała, o wyraźnie określonym kształcie. Cumulus oznacza po łacinie „sterta”, odnosząc się do charakterystycznego wyglądu tych chmur. Ale cumulus ma kilka różnych etapów ewolucji...


Fakty, które mogą nauczyć cię czegos nowego
Świat jest pełen fascynujących faktów i wiedzy czekających na odkrycie, z których każdy ma potencjał poszerzania naszych horyzontów i zmiany perspektyw.


Rzeczy, które zmieniły świat na gorszy
Technologia nie stoi w miejscu, tylko ciągle się rozwija. Jednak czasami rozwój cywilizacji jest okupiony działaniami, które zdecydowanie zmierzają w złą stronę...


Szalona historia życia Johna McAfee, miliardera i twórcy programu antywirusowego McAfee
23 czerwca 2021 roku w celi jednego z hiszpańskich więzień powiesił się osadzony John McAfee. Wiadomość o jego śmierci została opublikowana przez najważniejsze gazety i portale informacyjne. Denat to oczywiście nikt inny jak skandalizujący miliarder i programista, który kiedyś stworzył popularny program antywirusowy McAfee. Jednak w ostatnich latach jego nazwisko częściej pojawiało się w raportach kryminalnych niż w opracowaniach IT.


Skąd wziął się kolor pomarańczowy?
Może na pierwszy rzut oka tytułowe pytanie brzmi dziwnie, ale zastanawialiście się kiedyś, skąd się wziął kolor pomarańczowy? I mamy tu na myśli słowo, ale również sam kolor, barwniki. Czy jest w nim coś wyjątkowego? Okazuje się, że owszem.


Niezwykłe spojrzenie na zwykłe rzeczy artysty Redmera Hoekstry
Bujna wyobraźnia jest niezbędną cechą każdego artysty. Holenderski ilustrator i grafik Redmer Hoekstra także jest wizjonerem. Przedstawia znane stworzenia i najzwyklejsze przedmioty w taki sposób, że zmienia to nasze o nich wyobrażenie.
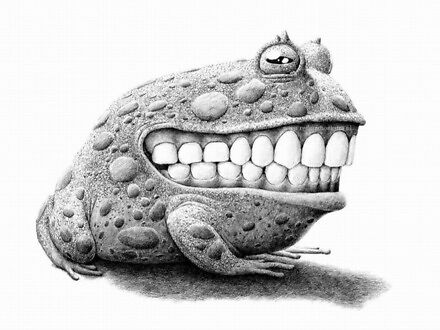


I weź tu sprzedaj lustro bez ośmieszania się przed całym internetem
Naprawdę mocno próbowali. Ale nie wyszło. Znaczy wyszło jak zawsze, a my możemy się pośmiać.


Azjatki, które mają w sobie to coś
Zdjęcia pięknych i seksownych mieszkanek Dalekiego Wschodu i Azji przenoszą nas w świat subtelnego wdzięku i egzotycznej fascynacji. Te uchwycone chwile nie tylko ukazują urodę kobiet, ale także emanują tajemniczym urokiem i delikatnością. Kulturowe dziedzictwo i tradycje stają się tu nie tylko elementem kontekstu, lecz również dodatkowym źródłem inspiracji dla ich wyjątkowej urody.


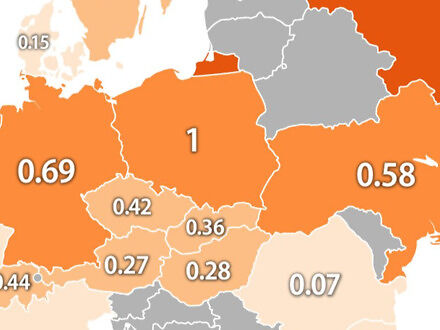
Przeprowadzono badanie, w którym poproszono uczestników, aby narysowali z pamięci logo znanych marek samochodów
Pewna brytyjska firma zajmująca się sprzedażą używanych samochodów postanowiła sprawdzić, jak dobrze jesteśmy w stanie zapamiętać logo znanych marek samochodów. Przeprowadzono więc mały eksperyment i poproszono każdego uczestnika, aby z pamięci spróbował narysować najbardziej rozpoznawalne znaczki w branży motoryzacyjnej. Oto rezultaty.


9 wyjątkowo głupich sposobów, w jakie ludzie przypadkowo pozbawili się życia
Pech, głupota, skrajny debilizm? Nazwijcie to sami!
#1. Isadora Duncan i rosyjski szal śmierci

7 ciekawostek o Comic Sans - najbardziej znienawidzonym foncie świata
Zupełnie niewinny font zdobył znienacka ogromną popularność. I jak szybko stał się sławny, tak szybko stał się też wrogiem trzech czwartych skomputeryzowanego świata.


Mistrzowie Internetu – Dlaczego w Polsce mamy wschodnie zarobki i zachodnie ceny
W dzisiejszym odcinku m.in. jak dostać się do piwa w tym sklepie; polski program kosmiczny; ile istnieje płci; kariery dzieci komunistycznych dygnitarzy w polskiej TV; nowa polska wojenka o alkohol na stacjach oraz jeśli ci smutno, to pomyśl o tym chłopaku w przedstawieniu i jego roli.


Za mocno kichnąłem i coś strzeliło mi w plecach – Ludzie, którzy mieli bardzo zły dzień
Pechowi ludzie tworzą tę pechową serię. Jeśli w życiu Ci nie idzie, to zobacz, że inni mają jeszcze gorzej.


Najmocniejsze cytaty – Ryszard Petru tłumaczy, że bez sprzedaży alkoholu stacje benzynowe nie przetrwają
Dzisiaj:

- Andrzej Duda powiedział, co myśli o Donaldzie Trumpie po ich spotkaniu
- Obajtek żali się, że jest śledzony i inwigilowany
- Nowy burmistrz Zakopanego zapowiada, że to koniec z Sylwestrem Marzeń
- Ryszard Petru tłumaczy, że bez sprzedaży alkoholu stacje benzynowe nie przetrwają
- Olechowski alarmuje! Nowe stroje polskich olimpijczyków przypominają flagę Rosji

Stopień zniszczenia niemieckich miast w czasie II wojny światowej – Kolekcja intrygujących map
W dzisiejszym odcinku m.in. 25 najstarszych demokracji na świecie, obecność wojskowa państw spoza Afryki na terenie tego kontynentu oraz stopień zniszczenia niemieckich miast w czasie II wojny światowej.

| Z archiwów JM |
|
|
Monster Galeria: Prawda jest... niewygodna
Najpotworniejsze ostatnio
Login
Nie masz jeszcze konta na tej najlepszej na świecie stronie?! Jakże tak można?!
Jako zarejestrowany bojownik JM, będziesz miał parę bonusów:
- strona będzie cie witać
- zmienisz sobie wygląd strony
- od czasu do czasu dostaniesz od nas maila
- będziesz mógł popisać się na forum
- będziesz recenzentem i oceniaczem
- skomentujesz nas szczerze
- obejrzysz w całości słynną MonsterGalerię, teraz to 310 257 obrazków. I wiem, że zawsze chciałeś ją obejrzeć…
- włączysz sobie na pełen regulator Szafę Grającą, teraz ponad 1564147 pliczków.
Krótkie dżołki
Nie udał się strajk listonoszy Poczty Polskiej. Okazuje się, że zamiast postulatów zostawili awizo.
* * *
Sztuczne ognie dla gamoni: frajerwerki.
* * *
Film był tak nudny, że na sali całowali się nawet nieznajomi ludzie.
* * *
Przedwczoraj był Dzień Książki. Postanowiłem go uczcić. Jestem już na trzeciej stronie.
* * *
Patrycja tak bardzo powiększyła sobie wargi, że faceci przestali zwracać uwagę na wąsy.
* * *
* * *
Sztuczne ognie dla gamoni: frajerwerki.
* * *
Film był tak nudny, że na sali całowali się nawet nieznajomi ludzie.
* * *
Przedwczoraj był Dzień Książki. Postanowiłem go uczcić. Jestem już na trzeciej stronie.
* * *
Patrycja tak bardzo powiększyła sobie wargi, że faceci przestali zwracać uwagę na wąsy.
* * *
Ciekawe wątki
Zakazane strony
Najlepsze komentarze
 „@puhacz Może gdyby nie robili antysemity z każdego kto się na nich krzywo popatrzy podejście byłoby ździebko inne...”
by geniusm
530
„@puhacz Może gdyby nie robili antysemity z każdego kto się na nich krzywo popatrzy podejście byłoby ździebko inne...”
by geniusm
530
 „Dobrze, że absolutnie każdy jest ekspertem od wszystkiego, ma dostęp do pełnych informacji, w tym niejawnych, i ma kompetencje do prowadzenia polityki zagranicznej/wstaw aktualnie nośny medialnie temat.
I ja nie twierdzę, że mamy popierać Izrael, tylko, że ten tok rozumowania przedstawiony na obrazku jest totalnie z dupy.”
by ka2
452
„Dobrze, że absolutnie każdy jest ekspertem od wszystkiego, ma dostęp do pełnych informacji, w tym niejawnych, i ma kompetencje do prowadzenia polityki zagranicznej/wstaw aktualnie nośny medialnie temat.
I ja nie twierdzę, że mamy popierać Izrael, tylko, że ten tok rozumowania przedstawiony na obrazku jest totalnie z dupy.”
by ka2
452
 „Można, ale pod warunkiem, że masz wujka w policji...”
by skaba
426
„Można, ale pod warunkiem, że masz wujka w policji...”
by skaba
426
 „Widzę to oczami wyobraźni i mi się podoba :)”
by bartoslaw
416
„Widzę to oczami wyobraźni i mi się podoba :)”
by bartoslaw
416
 „Nawet nie chce wiedzieć kto to jest :)”
by blinkovsky
405
„Nawet nie chce wiedzieć kto to jest :)”
by blinkovsky
405
Sprawdź swoją wiedzę!